(Re)edited: 2025-10-23
Load data
This is a departure from the previous installments, as we are loading in a very processed dataset. The reasons why are numerous:
The original data is 330mb, compressed
After loading (2 minutes on my quite fast computer) and uncompressing, it takes over 10GB of RAM on my computer
The original data needed a severe amount of massaging to make it quickly usable:
Annotating with gene name
Identifying primary transcript for gene
Expanding range data into row-form*
mosdepth provides read depth as ranges. So instead of writing a row for each of the three billion+ base pairs, mosdepth collapses adjacent rows with identical read depth together. This is crucial for saving space, especially for exomes, which have huge stretches of zero coverage since only ~3% of the genome is targeted.
Curious?
If you want to see how I did the above, see Part 2 . This post also has some cooler plots.
Data
dd_class.csv can be found here
── Attaching packages ─────────────────────────────────────── tidyverse 1.3.2 ──
✔ ggplot2 3.4.0 ✔ purrr 1.0.1
✔ tibble 3.1.8 ✔ dplyr 1.1.0
✔ tidyr 1.3.0 ✔ stringr 1.5.0
✔ readr 2.1.3 ✔ forcats 1.0.0
── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()
Attaching package: 'data.table'
The following objects are masked from 'package:dplyr':
between, first, last
The following object is masked from 'package:purrr':
transpose
library (cowplot) # you may need to install this with install.packages('cowplot') <- fread ('~/git/Let_us_plot/003_coverage/dd_class.csv' )head (dd_class)
Transcript Exon Number Start Chr End Read_Depth Strand
1: ENST00000020945.3 0 49831356 8 49831367 108 -
2: ENST00000020945.3 0 49831357 8 49831367 108 -
3: ENST00000020945.3 0 49831358 8 49831367 108 -
4: ENST00000020945.3 0 49831359 8 49831367 108 -
5: ENST00000020945.3 0 49831360 8 49831367 108 -
6: ENST00000020945.3 0 49831361 8 49831367 108 -
ExonStart ExonEnd Name
1: 49831365 49831547 SNAI2
2: 49831365 49831547 SNAI2
3: 49831365 49831547 SNAI2
4: 49831365 49831547 SNAI2
5: 49831365 49831547 SNAI2
6: 49831365 49831547 SNAI2
How many genes are in this dataset?
$ Name %>% unique () %>% length ()
What genes are in here?
$ Name %>% unique () %>% sort ()
[1] "ABCA4" "ABCB6" "AIPL1" "ALDH1A3" "ARL6" "ATF6"
[7] "ATOH7" "BBIP1" "BBS1" "BBS10" "BBS12" "BBS5"
[13] "BBS7" "BBS9" "BCOR" "BMP4" "CACNA1F" "CACNA2D4"
[19] "CASK" "CDHR1" "CEP290" "CHD7" "CNGB3" "CNNM4"
[25] "COL4A1" "COX7B" "CRX" "CYP1B1" "DCDC1" "EDN3"
[31] "EDNRB" "ELP4" "FKRP" "FKTN" "FOXC1" "FOXC2"
[37] "FOXE3" "GDF3" "GDF6" "GNAT2" "HESX1" "HMGB3"
[43] "HPS1" "HPS3" "HPS4" "HPS5" "HPS6" "INPP5E"
[49] "ISPD" "KCNJ13" "KCNV2" "KIT" "LAMB2" "LHX2"
[55] "LYST" "LZTFL1" "MAB21L2" "MFRP" "MITF" "MKKS"
[61] "MKS1" "MLPH" "MYO5A" "NAA10" "NDP" "NPHP1"
[67] "NRL" "OCA2" "OTX2" "PAX2" "PAX3" "PAX6"
[73] "PDE6C" "PDE6H" "PITPNM3" "PITX2" "PITX3" "POMT1"
[79] "POMT2" "PROM1" "PRPH2" "PRSS56" "RAB18" "RAB27A"
[85] "RAB3GAP1" "RAB3GAP2" "RARB" "RAX" "RAX2" "RDH5"
[91] "RET" "RIMS1" "RPGR" "RPGRIP1" "SDCCAG8" "SEMA4A"
[97] "SHH" "SIX3" "SIX6" "SLC24A5" "SLC25A1" "SLC38A8"
[103] "SLC45A2" "SNAI2" "SNX3" "SOX10" "SOX2" "SOX3"
[109] "STRA6" "TENM3" "TMEM98" "TRIM32" "TTC8" "TTLL5"
[115] "UNC119" "VAX1" "VSX2" "ZEB2"
How many data points (bases) per gene?
%>% group_by (Name) %>% summarise (Count= n ())
# A tibble: 118 × 2
Name Count
<chr> <int>
1 ABCA4 6804
2 ABCB6 2527
3 AIPL1 1159
4 ALDH1A3 1550
5 ARL6 569
6 ATF6 2008
7 ATOH7 457
8 BBIP1 220
9 BBS1 1773
10 BBS10 2172
# … with 108 more rows
How many exons per gene?
%>% select (Name, ` Exon Number ` ) %>% unique () %>% group_by (Name) %>% summarise (Count = n ())
# A tibble: 118 × 2
Name Count
<chr> <int>
1 ABCA4 50
2 ABCB6 19
3 AIPL1 6
4 ALDH1A3 13
5 ARL6 7
6 ATF6 16
7 ATOH7 1
8 BBIP1 2
9 BBS1 17
10 BBS10 2
# … with 108 more rows
How many base pairs of ABCA4 (well, ABCA4 exons) is covered by more than 10 reads?
Base R style
# Grab the Read_Depth vector from the data frame filtered by ABCA4 values <- dd_class %>% filter (Name== 'ABCA4' ) %>% pull (Read_Depth)sum (depth_abca4 > 10 )
Let’s check all of the genes to see which are the worst covered
%>% group_by (Name) %>% summarise (Total_Bases = n (),LT5 = sum (Read_Depth < 5 ),LT10 = sum (Read_Depth < 10 ),Good = sum (Read_Depth >= 10 ),P5 = LT5 / Total_Bases,P10 = LT10 / Total_Bases) %>% arrange (- P10)
# A tibble: 118 × 7
Name Total_Bases LT5 LT10 Good P5 P10
<chr> <int> <int> <int> <int> <dbl> <dbl>
1 BBIP1 220 0 61 159 0 0.277
2 ISPD 1339 0 57 1282 0 0.0426
3 CASK 2778 0 79 2699 0 0.0284
4 NAA10 725 0 10 715 0 0.0138
5 CNGB3 2436 0 18 2418 0 0.00739
6 LYST 11400 13 66 11334 0.00114 0.00579
7 PROM1 2601 0 8 2593 0 0.00308
8 RET 3348 0 9 3339 0 0.00269
9 CEP290 7456 0 18 7438 0 0.00241
10 SLC25A1 933 0 2 931 0 0.00214
# … with 108 more rows
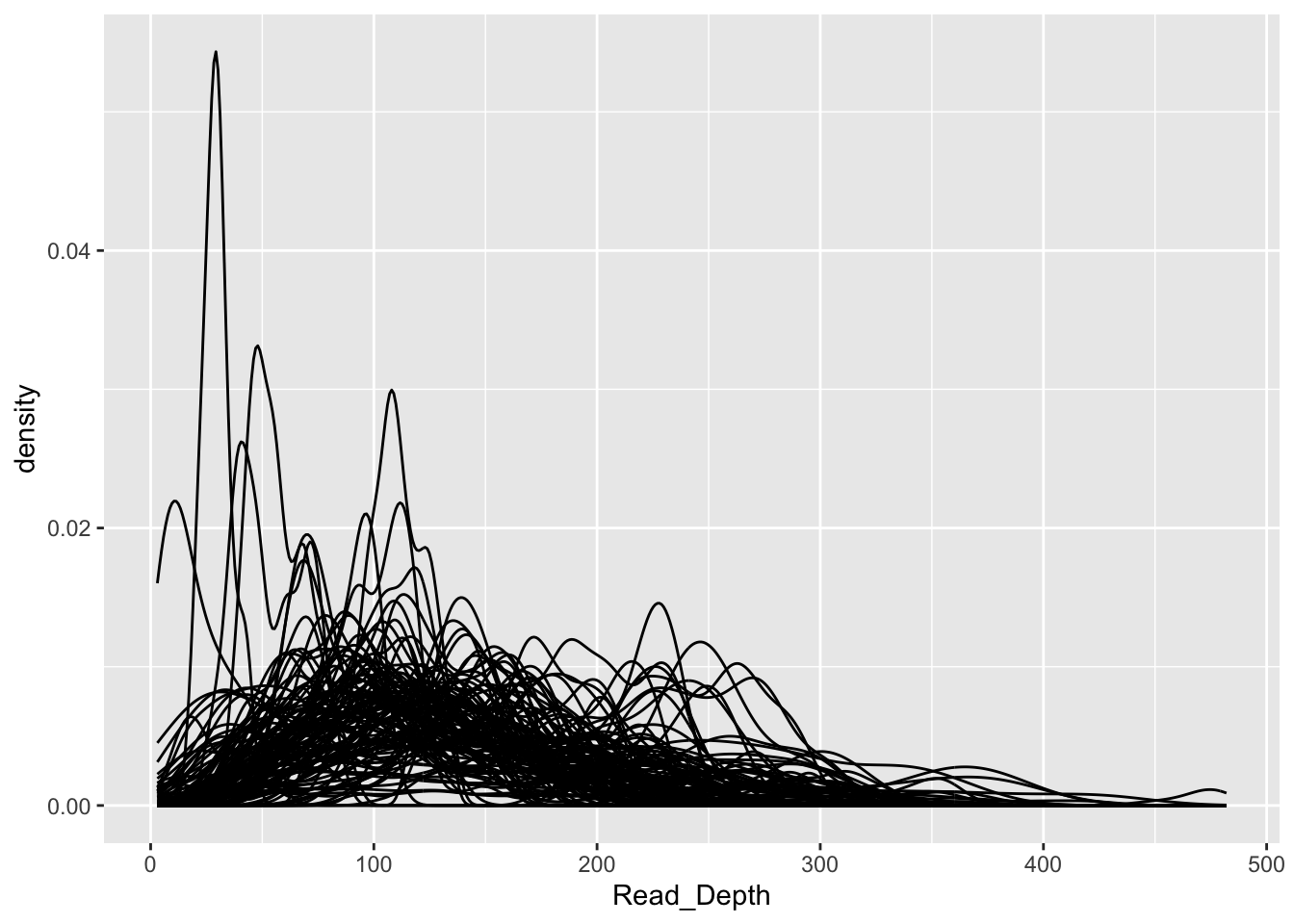
We can visually display the data, also
%>% ggplot (aes (x= Read_Depth, group= Name)) + geom_density ()
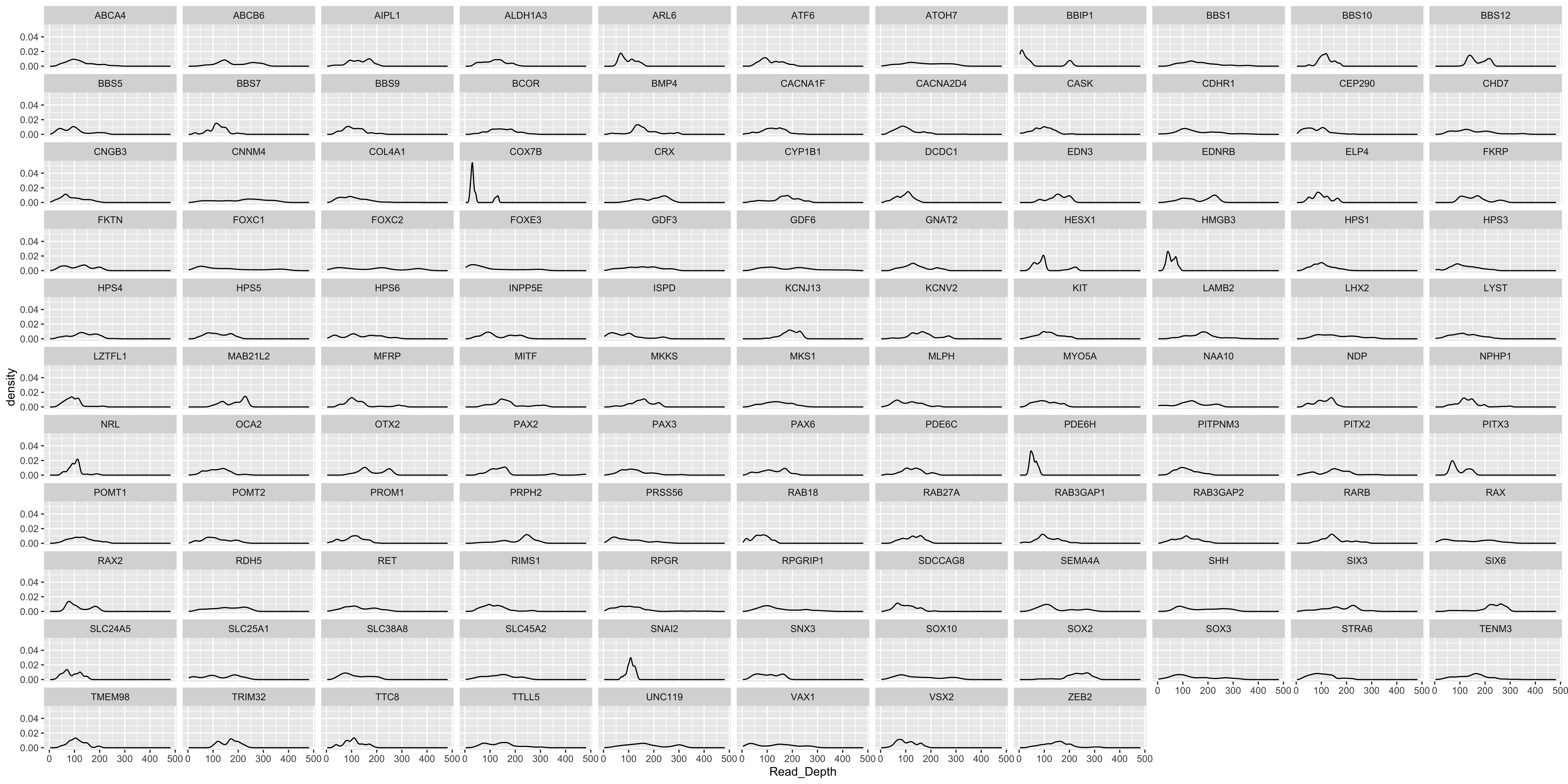
Hard to see what is going on, let’s make little plots for each gene
%>% ggplot (aes (x= Read_Depth, group= Name)) + facet_wrap (~ Name) + geom_density ()
##
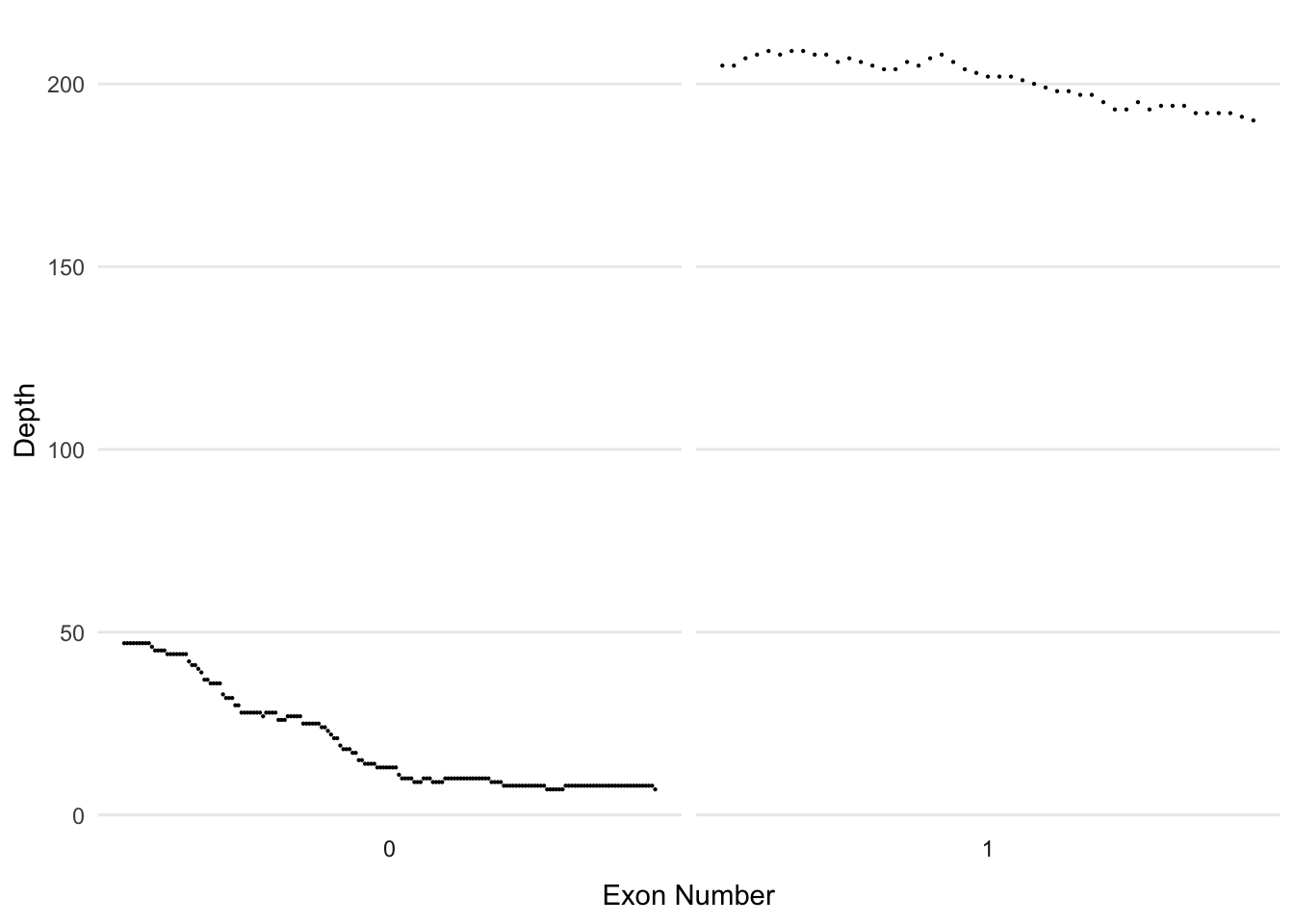
Where are genes poorly covered?
BBIP1
%>% filter (Name== 'BBIP1' ) %>% ggplot (aes (x= Start, y= Read_Depth)) + facet_wrap (~ ` Exon Number ` , scales = 'free_x' , nrow= 1 , strip.position = 'bottom' ) + geom_point (size= 0.1 ) + theme_minimal () + theme (axis.text.x= element_blank (), axis.ticks.x = element_blank (), panel.grid.minor = element_blank (), panel.grid.major.x = element_blank (),legend.position = 'none' ) + ylab ('Depth' ) + xlab ('Exon Number' )
Make a coverage plot for many genes (This is advanced stuff!!!)
<- c ('ABCA4' , 'PITX2' ,'VSX2' ,'RPGR' ,'SOX10' )# set a custom color that will work even if a category is missing <- function (...){::: manual_scale ('colour' , values = setNames (c ('darkred' , 'red' , 'black' ),c ('< 10 Reads' ,'< 20 Reads' ,'>= 20 Reads' )), <- function (gene){<- dd_class%>% filter (Name== gene) %>% pull (` Exon Number ` ) %>% as.numeric () %>% max ()# expand to create a row for each sequence and fill in previous values %>% filter (Name== gene) %>% mutate (` Exon Number ` = factor (` Exon Number ` ,levels= 0 : num_of_exons)) %>% ggplot (aes (x= Start, y= Read_Depth)) + facet_wrap (~ ` Exon Number ` , scales = 'free_x' , nrow= 1 , strip.position = 'bottom' ) + geom_point (size= 0.1 ) + theme_minimal () + scale_colour_custom () + theme (axis.text.x= element_blank (), axis.ticks.x = element_blank (), panel.grid.minor = element_blank (), panel.grid.major.x = element_blank (),legend.position = 'none' ) + ylab ('Depth' ) + xlab (gene)<- list ()for (i in gene_list){<- plot_maker (i)plot_grid (plotlist = plots, ncol= 1 )
─ Session info ───────────────────────────────────────────────────────────────
setting value
version R version 4.2.2 (2022-10-31)
os macOS 14.7.4
system aarch64, darwin20
ui X11
language (EN)
collate en_US.UTF-8
ctype en_US.UTF-8
tz America/New_York
date 2025-10-23
pandoc 3.6.3 @ /Applications/RStudio.app/Contents/Resources/app/quarto/bin/tools/aarch64/ (via rmarkdown)
─ Packages ───────────────────────────────────────────────────────────────────
package * version date (UTC) lib source
assertthat 0.2.1 2019-03-21 [1] CRAN (R 4.2.0)
backports 1.4.1 2021-12-13 [1] CRAN (R 4.2.0)
broom 1.0.3 2023-01-25 [1] CRAN (R 4.2.0)
cachem 1.0.6 2021-08-19 [1] CRAN (R 4.2.0)
callr 3.7.3 2022-11-02 [1] CRAN (R 4.2.0)
cellranger 1.1.0 2016-07-27 [1] CRAN (R 4.2.0)
cli 3.6.0 2023-01-09 [1] CRAN (R 4.2.0)
colorspace 2.1-0 2023-01-23 [1] CRAN (R 4.2.0)
cowplot * 1.1.1 2020-12-30 [1] CRAN (R 4.2.0)
crayon 1.5.2 2022-09-29 [1] CRAN (R 4.2.0)
data.table * 1.14.6 2022-11-16 [1] CRAN (R 4.2.0)
DBI 1.1.3 2022-06-18 [1] CRAN (R 4.2.0)
dbplyr 2.3.0 2023-01-16 [1] CRAN (R 4.2.0)
devtools 2.4.5 2022-10-11 [1] CRAN (R 4.2.0)
digest 0.6.31 2022-12-11 [1] CRAN (R 4.2.0)
dplyr * 1.1.0 2023-01-29 [1] CRAN (R 4.2.0)
ellipsis 0.3.2 2021-04-29 [1] CRAN (R 4.2.0)
evaluate 0.20 2023-01-17 [1] CRAN (R 4.2.0)
fansi 1.0.4 2023-01-22 [1] CRAN (R 4.2.0)
farver 2.1.1 2022-07-06 [1] CRAN (R 4.2.0)
fastmap 1.1.0 2021-01-25 [1] CRAN (R 4.2.0)
forcats * 1.0.0 2023-01-29 [1] CRAN (R 4.2.0)
fs 1.6.1 2023-02-06 [1] CRAN (R 4.2.0)
gargle 1.3.0 2023-01-30 [1] CRAN (R 4.2.0)
generics 0.1.3 2022-07-05 [1] CRAN (R 4.2.0)
ggplot2 * 3.4.0 2022-11-04 [1] CRAN (R 4.2.0)
glue 1.6.2 2022-02-24 [1] CRAN (R 4.2.0)
googledrive 2.0.0 2021-07-08 [1] CRAN (R 4.2.0)
googlesheets4 1.0.1 2022-08-13 [1] CRAN (R 4.2.0)
gtable 0.3.1 2022-09-01 [1] CRAN (R 4.2.0)
haven 2.5.1 2022-08-22 [1] CRAN (R 4.2.0)
hms 1.1.2 2022-08-19 [1] CRAN (R 4.2.0)
htmltools 0.5.4 2022-12-07 [1] CRAN (R 4.2.0)
htmlwidgets 1.6.1 2023-01-07 [1] CRAN (R 4.2.0)
httpuv 1.6.9 2023-02-14 [1] CRAN (R 4.2.0)
httr 1.4.4 2022-08-17 [1] CRAN (R 4.2.0)
jsonlite 1.8.4 2022-12-06 [1] CRAN (R 4.2.0)
knitr 1.42 2023-01-25 [1] CRAN (R 4.2.0)
labeling 0.4.2 2020-10-20 [1] CRAN (R 4.2.0)
later 1.3.0 2021-08-18 [1] CRAN (R 4.2.0)
lifecycle 1.0.3 2022-10-07 [1] CRAN (R 4.2.0)
lubridate 1.9.1 2023-01-24 [1] CRAN (R 4.2.0)
magrittr 2.0.3 2022-03-30 [1] CRAN (R 4.2.0)
memoise 2.0.1 2021-11-26 [1] CRAN (R 4.2.0)
mime 0.12 2021-09-28 [1] CRAN (R 4.2.0)
miniUI 0.1.1.1 2018-05-18 [1] CRAN (R 4.2.0)
modelr 0.1.10 2022-11-11 [1] CRAN (R 4.2.0)
munsell 0.5.0 2018-06-12 [1] CRAN (R 4.2.0)
pillar 1.8.1 2022-08-19 [1] CRAN (R 4.2.0)
pkgbuild 1.4.0 2022-11-27 [1] CRAN (R 4.2.0)
pkgconfig 2.0.3 2019-09-22 [1] CRAN (R 4.2.0)
pkgload 1.3.2 2022-11-16 [1] CRAN (R 4.2.0)
prettyunits 1.1.1 2020-01-24 [1] CRAN (R 4.2.0)
processx 3.8.0 2022-10-26 [1] CRAN (R 4.2.0)
profvis 0.3.7 2020-11-02 [1] CRAN (R 4.2.0)
promises 1.2.0.1 2021-02-11 [1] CRAN (R 4.2.0)
ps 1.7.2 2022-10-26 [1] CRAN (R 4.2.0)
purrr * 1.0.1 2023-01-10 [1] CRAN (R 4.2.0)
R6 2.5.1 2021-08-19 [1] CRAN (R 4.2.0)
Rcpp 1.0.10 2023-01-22 [1] CRAN (R 4.2.0)
readr * 2.1.3 2022-10-01 [1] CRAN (R 4.2.0)
readxl 1.4.1 2022-08-17 [1] CRAN (R 4.2.0)
remotes 2.4.2 2021-11-30 [1] CRAN (R 4.2.0)
reprex 2.0.2 2022-08-17 [1] CRAN (R 4.2.0)
rlang 1.0.6 2022-09-24 [1] CRAN (R 4.2.0)
rmarkdown 2.20 2023-01-19 [1] CRAN (R 4.2.0)
rstudioapi 0.14 2022-08-22 [1] CRAN (R 4.2.0)
rvest 1.0.3 2022-08-19 [1] CRAN (R 4.2.0)
scales 1.2.1 2022-08-20 [1] CRAN (R 4.2.0)
sessioninfo 1.2.2 2021-12-06 [1] CRAN (R 4.2.0)
shiny 1.7.4 2022-12-15 [1] CRAN (R 4.2.0)
stringi 1.7.12 2023-01-11 [1] CRAN (R 4.2.0)
stringr * 1.5.0 2022-12-02 [1] CRAN (R 4.2.0)
tibble * 3.1.8 2022-07-22 [1] CRAN (R 4.2.0)
tidyr * 1.3.0 2023-01-24 [1] CRAN (R 4.2.0)
tidyselect 1.2.0 2022-10-10 [1] CRAN (R 4.2.0)
tidyverse * 1.3.2 2022-07-18 [1] CRAN (R 4.2.0)
timechange 0.2.0 2023-01-11 [1] CRAN (R 4.2.0)
tzdb 0.3.0 2022-03-28 [1] CRAN (R 4.2.0)
urlchecker 1.0.1 2021-11-30 [1] CRAN (R 4.2.0)
usethis 2.1.6 2022-05-25 [1] CRAN (R 4.2.0)
utf8 1.2.3 2023-01-31 [1] CRAN (R 4.2.0)
vctrs 0.5.2 2023-01-23 [1] CRAN (R 4.2.0)
withr 2.5.0 2022-03-03 [1] CRAN (R 4.2.0)
xfun 0.37 2023-01-31 [1] CRAN (R 4.2.0)
xml2 1.3.3 2021-11-30 [1] CRAN (R 4.2.0)
xtable 1.8-4 2019-04-21 [1] CRAN (R 4.2.0)
yaml 2.3.7 2023-01-23 [1] CRAN (R 4.2.0)
[1] /Library/Frameworks/R.framework/Versions/4.2-arm64/Resources/library
──────────────────────────────────────────────────────────────────────────────